SARS coronavirus is a virus that causes severe acute respiratory syndrome (SARS). On April 16, 2003, following the outbreak of SARS in Asia and secondary cases elsewhere in the world, the World Health Organization (WHO) issued a press release stating that the coronavirus identified by a number of laboratories was the official cause of SARS. Samples of the virus are being held in laboratories in New York, San Francisco, Manila, Hong Kong, and Toronto. It emerged from 'somewhere' and lead to many problems like; 8000 people infected and killed nearly 700 people.
• Known viruses that has newly appeared in a human population and is rapidly increasing in disease incidence.
• Spontaneous evolution of new virus entity.
- Many viruses, in particular RNA viruses, have short generation times and relatively high mutation rates (on the order of one point mutation or more per genome per round of replication for RNA viruses). This elevated mutation rate, when combined with natural selection, allows viruses to quickly adapt to changes in their host environment.
• Generation of a novel strain due to co-infection of different strains in an individual (random assortment)
- Many viruses, in particular RNA viruses, have short generation times and relatively high mutation rates (on the order of one point mutation or more per genome per round of replication for RNA viruses). This elevated mutation rate, when combined with natural selection, allows viruses to quickly adapt to changes in their host environment.
- When the underlying causes and mechanisms of emerging infectious disease problems are studied carefully, human behavior is often involved. Even more often, the only methods of control or prevention available are to change human behavior. Several major recent emerging disease problems can be cited. It is sometimes emphasized that it is human carelessness, human excesses, human ignorance or human habits of conquest or leisure which contribute directly to the biological niches that microorganisms are all too capable of exploiting.
- Climate change is increasingly becoming a concern as a factor in the emergence of infectious diseases. As Earth's climate warms and habitats are altered, diseases can spread into new geographic areas. In fact, this has already happened. For the first time, a tropical disease has caused an outbreak in Europe. In late summer of 2007, more than 100 residents of the town of Ravenna, Italy suffered from a mysterious disease that produced fever, exhaustion, and severe bone pain. The outbreak was eventually shown to be caused by chikungunya virus, a relative of the virus that causes Dengue fever, previously found in tropical regions around the Indian Ocean. Due to warming and globalization, the tiger mosquito which transmits chikungunya virus has been able to move north and thrive in areas across southern Europe and spread the chikungunya virus. Although chikungunya virus does not usually cause a fatal disease, this outbreak serves as a warning that other, more devastating tropical diseases could follow.
- Many emerging diseases arise when infectious agents in animals are passed to humans (referred to as zoonoses). As the human population expands in number and into new geographical regions, the possibility that humans will come into close contact with animal species that are potential hosts of an infectious agent increases. When that factor is combined with increases in human density and mobility, it is easy to see that this combination poses a serious threat to human health.
Viral populations are heterogeneous. One viral genome obtained from an infected person may not be identical to genome from all the viruses. This is due to its high mutation rates like selection pressure.
The orientations of the homologous chromosomes at the equatorial plate of the spindle are random. That is, each time they line up at the equatorial plate, the two pairs of chromosomes in a homologue may appear on different sides of the plate. As you have already learned in the previous page, the alleles on the two homologous chromosomes are different. Thus, when the cell separates during meiosis, the resulting cells contain different genetic codes. This is called random assortment.
For instance, H3N2 infects man, H5N1 infect birds. Co-infection of both H3N2 and H5N1 means that they infect the same host together at the same time. The virus will replicate inside the host.
Only 4 strains of avian influenza virus are known to cause disease in humans; H5N1, H7N3, H7N7, H9N2.
• H5N1 is a subtype of the Influenza A virus which can cause illness in humans and many other animal species.
• H7N3 is a subtype of the species Influenza A virus (sometimes called bird flu virus).
• In North America, the presence of H7N3 was confirmed at several poultry farms in British Columbia in February 2004. As of April 2004, 18 farms had been quarantined to halt the spread of the virus. Two cases of humans infected with it have been confirmed in that region. Symptoms included conjunctivitis and mild influenza-like illness.
• H7N7 is a subtype of Influenzavirus A, a genus of Orthomyxovirus, the viruses responsible for influenza. Highly pathogenic strains (HPAI) and low pathogenic strains (LPAI) exist. H7N7 can infect humans, birds, pigs, seals, and horses in the wild; and has infected mice in laboratory studies. This unusual zoonotic potential represents a pandemic threat.
• H9N2 is a subtype of the species Influenza A virus (sometimes called bird flu virus). In 1999 and 2003, an H9N2 influenza strain caused illness in three people, aged one, four and five years old, in Hong Kong. All three patients recovered. In 2007 an H9N2 influenza strain caused illness in a 9-month old baby in Hong Kong.
Research had shown that out of 422 cases, 95% of people in 2007 were infected by viruses through Sex. Other cases like sharing of needles during injection also can promote new viruses emerging too.
This is not likely to happen in developed countries as they have the technology and manpower to fight against new emerging viruses. In less developed countries, these are more likely to happen as they have no proper healthcare.
Climate changes can increase the cases of emerging viruses. This is a human factor can cause this indirectly. For instance, via driving, there would be increase of global warming and this can cause a change in climate. Moreover, these climate changes may cause an increase in raining seasons, which promote the breed of mosquitoes that may lead to an increase in dengue cases.
In another case, haze can cause new viruses to emerge too. Breathing in of ‘plant hormones’ blown by the haze may cause some symptoms in humans that are currently unknown. It may also have some long tern effect in human after long exposure.
Human perform deforestation for agricultural needs. Through hunting and poaching, this bring human to have a closer proximity to those wild animals. This may cause an increase possibility of zoonoses. Zoonoses means that the disease that infect animals can infect human now.
So, what are the new viruses that will emerge over the next few years? Well, we don’t know and it’s best for us to prepare to fight against it now! That the end of the chapter on emerging viruses, next we’ll find out more about Prions, Viroids and Virusoids.
Prions are unprecedented infectious pathogens that cause a group of invariably fatal neurodegenerative diseases by an entirely novel mechanism. Prion diseases may present as genetic, infectious, or sporadic disorders, all of which involve modification of the prion protein (PrP). Bovine spongiform encephalopathy (BSE) of cow, scrapie of sheep, and Creutzfeldt-Jakob disease (CJD) of humans are among the most notable prion diseases. Prions are transmissible particles that are devoid of nucleic acid and seem to be composed exclusively of a modified protein (PrPSc).
Prions are rogue protein that transforms other cellular protein (PrPc) to the prion form PrPsc. PrPc gene is on chromosome 20 and more PrPsc gets transformed until they completely clogged brain cells thus causing cells to misfire, work poorly or don’t work at all. Cells will then die and release prions into blood stream to re-infect other cells. Prions are hypothesized to infect and propagate by refolding abnormally into a structure which is able to convert normal molecules of the protein into the abnormally structured form. All known prions induce the formation of an amyloid fold, in which the protein polymerises into an aggregate consisting of tightly packed beta sheets. This altered structure is extremely stable and accumulates in infected tissue, causing cell death and tissue damage. This stability means that prions are resistant to denaturation by chemical and physical agents, making disposal and containment of these particles difficult.
PrPC is a normal protein found on the membranes of cells. It has 209 amino acids (in humans), one disulfide bond, a molecular weight of 35-36kDa and a mainly alpha-helical structure. Several topological forms exist; one cell surface form anchored via glycolipid and two transmembrane forms. Its function has not been fully resolved. PrPC binds copper (II) ions with high affinity. The significance of this is not clear, but it presumably relates to PrP structure or function. PrPC is readily digested by proteinase K and can be liberated from the cell surface in vitro by the enzyme phosphoinositide phospholipase C (PI-PLC), which cleaves the glycophosphatidylinositol (GPI) glycolipid anchor.
The infectious isoform of PrPC, known as PrPSc, is able to convert normal PrPC proteins into the infectious isoform by changing their conformation, or shape; this, in turn, alters the way the proteins interconnect. Although the exact 3D structure of PrPSc is not known, infected cells show accumulation of β-sheet protein in place of the normal areas α-helix form. Aggregations of these abnormal isoforms form highly structured amyloid fibers, which accumulate to form plaques. The end of each fiber acts as a template onto which free protein molecules may attach, allowing the fiber to grow. Differences in the amino acid sequence of prion-forming regions lead to distinct structural features on the surface of the abnormal amyloid fibers. As a result, only free protein molecules identical to the prion protein are incorporated into the growing fiber.
Prion diseases are caused by conversion of a normally folded, non-pathogenic isoform of the prion protein PrPc to a misfolded, pathogenic isoform PrPsc. Prion inoculation experiments in mice expressing homologous PrPc molecules on different genetic backgrounds displayed different incubation times, indicating that the conversion reaction may be influenced by other gene products. To identify genes that contribute to prion pathogenesis, we analysed incubation times of prions in mice in which the gene product was inactivated, knocked out or over expressed.
PrPsc forms a heterodimer with normal PrPc. It is a template for altering the protein fold. It tightly coiled alpha helix converted to loose beta sheets. This was supported by using two mice in an experiment. One mouse contains knockout PrPc gene whereas the other contains normal PrPc gene. The mouse which contains knockout PrPc gene has no PrPc protein while the mouse which contains normal PrPc produces PrPc protein. Then, both mice are infected with prions. The mouse that contains knockout PrPc survives as there is nothing to convert the PrPc whereas the mouse that contains normal PrPc gene has prions to convert the PrPc to rogue proteins died.
Dementia, ataxia (loss of muscle control of voluntary movements) from the loss of brain functions due to degeneration.
In the past, diagnosis of prion disease was made through examination of brain biopsies taken from patients in advanced stages of the disease or, more commonly, after they had died. In January of 1999 it was found that the prion protein accumulated in the tonsils and could be detected by an immunofluorescence test on tonsilar biopsies. A second test was simultaneously developed which was based on a Western blot. Later that year a third test was developed that had the high sensitivity necessary to detect the prion protein in blood. This test is based on capillary electrophoresis with laser-induced fluorescence. It detects as little as 10-18 mole.
The primary method of prions infection in animals is through ingestion. It is thought that prions may be deposited in the environment through the remains of dead animals and via urine, saliva, and other body fluids. They may then linger in the soil by binding to clay and other minerals. The transmission of prions in humans is the consuming of contaminated beef or mutton, basically any type of contaminated meat.
Prion diseases are fatal neurodegenerative disorders of animals and humans that are characterized by the conversion of the host-encoded prion protein (PrP) to an abnormal isoform. In several species, including humans, polymorphisms in the gene encoding the PrP protein tightly control susceptibility of individuals toward this disease.
Viroids are infectious agents composed exclusively of a single piece of circular single stranded RNA which has some double-stranded regions. Viroid is a very small genome and in the case of prions, possibly no genome at all. Viroids affect mainly plants (25 main sequences identified) and only one sequence is known to infect man; which is Hepatitis D.
The RNA genomes of Viroids are 246-375 nucleotides in length and they share many similarities:
Catalytic RNAs are those that have the intrinsic ability to break and form covalent bonds; Viroids are catalytic RNA’s (ribozymes) that cleave RNA to produce fragments containing a 5’-hydroxl and a 2’, 3’-cyclic phosphate.
The only human disease known to be caused by a viroid is hepatitis D. This disease was previously ascribed to a defective virus called the delta agent. However, it now is known that the delta agent is a viroid enclosed in a hepatitis B virus capsid. For hepatitis D to occur there must be simultaneous infection of a cell with both the hepatitis B virus and the hepatitis D viroid. There are extensive sequence complementarities between the hepatitis D viroid RNA and human liver cell 7S RNA; a small cytoplasmic RNA that is a component of the signal recognition particle, the structure involved in the translocation of secretory and membrane-associated particles. The hepatitis D viroid causes liver cell death via sequestering this 7S RNA and/or cleaving it.
The hepatitis D viroid can only enter a human liver cell if it is enclosed in a capsid that contains a binding protein. It obtains this from the hepatitis B virus. The delta agent then enters the blood stream and can be transmitted via blood or serum transfusions.
Unlike viroids, virusoid is an infectious agent that infects plants in conjunction with an assistant virus. Virusoid is not considered a virus but a subviral particle. The size and structure is similar to viroids. Its genome is single molecule of single stranded circular RNA that is several hundred nucleotides long and codes for nothing but its own structure. Virusoids belong to a larger group of infectious agents called satellite RNAs, found in bacteria, plants, fungi, invertebrates and vertebrates. Satellite genomes encode proteins; satellite viruses encode capsid proteins but are still dependent upon a helper virus for replication.
Virusoid genomes are 220 to 388 nucleotides long. A virusoid genome does not code for any proteins, but instead serves only to replicate itself. Virusoids can replicate in the cytoplasm and possess a ribozyme activity. RNA replication is similar to that of viroids, but each requires that the cell be infected with a specific "helper" virus. Five virusoids are known, and the helper viruses for these are all members of the Sobemovirus family. An example of a "helper" virus is the subterranean clover mottle virus, which has an associated virusoid. Virus enzymes may aid replication of the virusoid RNA. The virusoid is incorporated into the virus particle and transmitted as a "satellite," a separate nucleic acid not part of the viral chromosome. Replication of the helper virus is independent of the virusoid.
Thats all for Lecture 9 and 10. We hope you enjoy reading our reviews on Virology! (:
LECTURE 7
Methods of Study of Viruses
Introduction to the Methods of Studying Viruses
The study of virus is Virology. The study of viruses is crucial to molecular and cellular biology as let us know the manipulation and functions of cells. Viruses are often used as a vector to introduce genes into the cells. After that, we would study the cell if it produces a new foreign substance or the effect of introducing a new gene into a genome.
Viruses are studied for isolation and cultivation of animals and plants cells, embryonated eggs and tissue culture.
Viruses can be detected, identified and diagnosed via various methods. For instances:
• Tissue culture methods
• Physical methods
• Serological methods
• Immunological methods
There are other methods available and molecular biology can be used too.
Animals, plants and eggs are the first method used for virus cultivation. Although it may be inconvenient, it is safer in handling animals. Most cultivations are replaced by cell culture except for those:
• Virus that has no known host in vitro
• The Hepatitis C virus in chimpanzees
• The influenza virus in chick embryo
The Hepatitis C virus in chimpanzees cannot be cultivated using cell culture as these viruses that recovered from animals are highly infectious in cell culture.
• Study of viral pathogenesis in a whole host.
• Polio studies in chimpanzees
Picture of Egg inoculation:
 In plants
In plants
• Tobacco mosaic virus
Tobacco mosaic virus is an RNA virus that infects plants, especially tobacco. This infection causes characteristic patterns like mottling and discoloration) on the leaves , thus the name mosaic in the disease. When Tobacco mosaic virus infects a tobacco plant, the virus enters mechanically and replicates. After its multiplication, it enters the neighboring cells. The first symptom of this virus disease is that light green colouration will appears between the veins of young leaves. Next, the development of a “mosaic” pattern of light and dark green areas in the leaves will show. This will not result in plant death, but if infection occurs early in the season, plants are stunted. Leaves that grow lower are subjected to “mosaic burn” especially during periods of hot and dry weather. In these cases, large dead areas develop in the leaves. This constitutes one of the most destructive phases of tobacco mosaic virus infection. Infected leaves may appear to be crinkled, puckered, or elongated.
Tobacco mosaic Plant:

• Virus plaques on leaves
Virus plaques are done on leaves to determine the number of viruses growing.
Tissue culture
Tissue culture is growth of cells separate from the organism. Through the use of a liquid, semi-solid, or solid growth media, such as broth or agar, cells are cultured. Tissue culture commonly refers to the culture of animal cells and tissues, while the more specific term plant tissue culture is used for plants. In this lecture, we will learn more about primary cell line and continuous cell lines.
The growth of explants taken directly from the living organism is also known as primary cell culture. Primary cell cultures are obtained directly from an animal and the differentiated state can keep for a short period (days to weeks). Functionally differentiated primary cell cultures have a limited life span, and although maintenance of the differentiated properties has been improved by additives to the culture medium, components of the extracellular matrix or by different forms of co-culture, cell specific functions will eventually decline.
Continuous cell line is a group of morphologically uniform cells that can be propagated in vitro for an indefinite time. These lines have an unlimited proliferation capacity and which originated from embryos, tumours or transformed cells.
Trypsinisation is a key stage in most cell culture protocols. In tissue cell culture, trypsin is used to re-suspend cells adherent to the cell culture dish wall during the process of harvesting cells primary cell line. Cell removal via trypsinisation requires constant vigilance to detect the point of cell detachment.
A Mouse Primary Cell line:
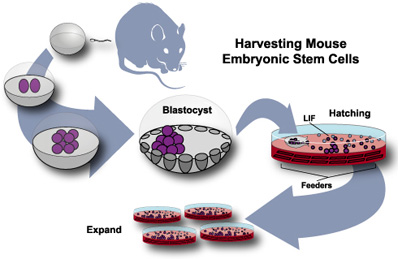
Trysinisation:
 Continuous Cell Line
Continuous Cell Line
Continuous cell line is:
• Derived from primary cell line
• Transformed/cancerous cells
• Polyploidy or multiploid
• Theoretically can be sub-cultured indefinitely
• Another method to cultivate virus
Tissue Culture Methods For Detection
There are 2 tissue culture methods for detection. They are cytopathic effect (CPE) and plaque assay.
Cytopathic effect (CPE) is the degenerative changes in cells (especially in tissue culture) associated with the multiplication of certain viruses; when, in tissue culture, spread of virus is restricted by an overlay of agar and the cytopathic effect may lead to formation of plaque.
CPE:

Plaque assay is developed by Renato Dulbecco in 1952; based on his original plaque assay for bacteriophage. It is used to observe cell death in infected cell culture; using the principle that only one virus infects one cell and spreads to the surrounding cells. To add on, it gives a higher accuracy at lower concentration as too many plaques would be labelled as too many to count (TNTC). Therefore, virus dilution is carried out to lower the concentration.
Plaque assay is developed to count and measure infectivity of bacteriophages and mammalian viruses. The plaque assay remains to be the most widely used technique for virus isolation and purification, and to determine viral titers. The basis of the technique is to measure the ability of a single infectious virus to form a plaque on a confluent monolayer culture of cells. A plaque is formed as a result of infection of one cell by a single virus particle followed by the replication of that virus, and eventually, cell lysis. The newly replicated virus particles will then infect and kill surrounding cells. The culture will then be stained with a trypan blue, which stains only viable cells but not the dead cells. Hence, the dead cells in the plaque will appear colourless against the coloured background.
The features of virus count are:
• Counts only viable cells as virus are capable of multiplying
• Must know the culture conditions of virus studied
• Useful for samples with very low virus counts as those with high virus count are TNTC
• Time required for incubation
Serial Dilution:

Plaque Assay:

Wells:
 Features Of Plaque Assay
Features Of Plaque Assay
The features for plaque assay are:
• very time-consuming
• very simple method
• only works for viruses that infect monolayer cells
• only works for viruses that cause cell lysis
• uses principle of one virus on confluent monolayer culture to produce one cell
Physical Methods
• X-ray crystallography
X-ray Crystallography is a method for solving the three dimensional structures of molecules at atomic resolution. It is used to determine the arrangement of atoms within a crystal, in which a beam of X-rays strikes a crystal and scatters into many different directions.
• Electron Microscopy
- Transmission Electron Microscope (TEM)
- Scanning Electron Microscope (SEM)
- Scanning Transmission Electron Microscope (STEM)
Electron Microscopes are scientific instruments that use a beam of highly energetic electrons to examine objects on a very fine scale. Transmission Electron Microscope (TEM) was the first type of Electron Microscope to be developed and is patterned exactly on the Light Transmission Microscope except that a focused beam of electrons is used instead of light to "see through" the specimen. Scanning Electron Microscope (SEM) does not at any time carry a complete image of the specimen. The SEM produces images by probing the specimen with a focused electron beam that is scanned across a rectangular area of the specimen. Scanning Transmission Electron Microscope (STEM) is a type of transmission electron microscope. The electrons in it pass through the specimen, but, as in scanning electron microscopy, the electron optics focuses the beam into a narrow spot which is scanned over the sample in a raster.
Pictures of Light and Electron Microscopes:

• Ultracentrifugation
• Purification
Purification is the process of rendering something pure.
Serological/Immunological Methods
Basic Immunology is an antigen.
• An antigen (from antibody-generating) is a substance that prompts the generation of antibodies and can cause an immune response in our body. Antigens are those substances that elicit a response from the immune system when foreign particle or molecule not recognized by the body is introduced.
• Examples are virus, bacteria, toxins, foreign particles and others.
• Hemagglutinin (HA)
• Hemagglutinin Inhibition (HI)
• Virus neutralization
• Complement fixation
• Immunoprecipitation/Immunoblot
• ELISA
• Immunostaining
- Immunoflourescene
- Immuno-gold EM
Hemagglutination
Hemagglutination is a specific form of agglutination that involves red blood cells. It has two common uses in the laboratory: blood typing and the quantification of virus dilutions. Examples are like influenza and other viruses. Influenza has two spike proteins like Hemagglutinin (HA) and Neuraminidase (NA). Hemagglutinin (HA) is responsible for binding the virus to the cell that is being infected. Neuraminidase (NA) is involved in the process of new virions budding out of host cells.
Picture of Hemagglutination:
 Hemagglutination Inhibition
Hemagglutination Inhibition
Hemagglutination Inhibition is Hemagglutinin (HA) conducted in the presence of anitibody. The neutralization of virus inhibits agglutination. Hemagglutination Inhibition measures of the ability of soluble antigen to inhibit the agglutination of antigen-coated red blood cells by antibodies. In this inhibition, a fixed amount of antibodies to the antigen is mixed with a fixed amount of red blood cells coated with the antigen.
Virus Neutralisation
Antibody neutralization prevents or lowers the virus infectivity. Virus infectivity is not blocked by an antibody that binds to a receptor for the virus on the cell surface. When high concentrations of antibody to the virus in question are present in the serum sample, virus neutralisation will occur even at high serum dilutions. Conversely, where little or no antibody to the virus is present in the test sample, it will be unable to neutralise the aliquot of infectious virus at the first dilution used in the test. The result of the test is the point at which the serum sample has been diluted such that it is no longer able to neutralise the entire virus in the test.
Complement Fixation
The complement fixation test is an immunological medical test that can be used to detect the presence of either specific antibody or specific antigen in a patient's serum. It was widely used to diagnose infections, particularly with microbes that are not easily detected by culture methods, and in rheumatic diseases. It is:
• Mediated by antibody
• Antibody binds to antigen
• Complement cascade of molecules in blood serum initiated, causing death in infected cell or pathogen.
Immunofluorescence
Immunofluorescence is a laboratory technique to identify specific antibodies or antigens. Antibody identification is usually performed on blood (serum). It is the labeling of antibodies or antigens with fluorescent dyes. This technique is often used to visualize the subcellular distribution of biomolecules of interest. Immunofluorescent-labelled tissue sections or cultures are studied using a fluorescence microscope or by confocal microscopy. In point form, Immunofluorescence is:
• Anitibody tagged with fluorescent dye
• Antibody attached specially to antigen
• Specimen is viewed under exciting light
• Through use of fluorescence microscope
Immunogold Electron Microscopy
Immunogold Electron Microscopy means that in a fluorescent microscope, antibodies with fluorescent markers are used to identify and locate specific macromolecules. An analogous method to this is the immunogold electron microscopy, in which a secondary antibody is tagged with a colloidal gold particle. This antibody is specific for a primary antibody that attaches to a macromolecule of interest. As the gold particle is electron dense, it will appear as a black dot on the TEM. This method will only detect tagged antibodies on the surface of the cut. In short, Immunogold Electron Microscopy uses:
• Sample principle as Immunofluorescence
• Gold particles attached to antibodies (nanometer size paticles)
• EM to view to localize specific proteins or antigens
Immunoprecipitation
Immunoprecipitation (IP) is the technique of precipitating a protein antigen out of solution using an antibody that specifically binds to that particular protein. This process can be used to isolate and concentrate a particular protein from a sample containing many thousands of different proteins. Immunoprecipitation requires that the antibody be coupled to a solid substrate at some point in the procedure. There are four types of Immunoprecipitation;
- Individual protein Immunoprecipitation (IP)
- Protein complex Immunoprecipitation (Co-IP)
- Chromatin Immunoprecipitation (ChIP)
- RNA Immunoprecipitation (RIP).
In short, Immunoprecipitation uses:
• Antigen radioactively labelled
• Reaction with antibody
• Isolate antibody-antigen complex
• Run through SDS-PAGE
• Detect through x-ray film
Immunoblot
Immunoblot is an analytical technique used to detect specific proteins in a given sample of tissue homogenate or extract. It uses gel electrophoresis to separate native or denatured proteins by the length of the polypeptide (denaturing conditions) or by the 3-D structure of the protein (native/ non-denaturing conditions). The proteins are then transferred to a membrane (typically nitrocellulose or PVDF), where they are probed (detected) using antibodies specific to the target protein. In summary, Immunoblot is:
• Western blot analysis
• Whole protein sample run through SDS-PAGE
• Transfer to nitrocellulose membrane
• Antibody attaches specifically to one protein
• Antibody labeled with sensitive indicator like horse radish peroxidase
• Colour reaction with streptavidin
A picture of Western Blot:
 ELISA
ELISA
Elisa is called Enzyme-Linked ImmunoSorbent Assay, or Enzyme ImmunoAssay or in short EIA. It is a biochemical technique used mainly in immunology to detect the presence of an antibody or an antigen in a sample. A type of enzyme immunoassay (EIA) is to determine the presence of antibodies to HIV in the blood or oral fluids. A reactive ELISA test results should be validated with an independent supplemental test of high specificity. There are 4 types of ELISA, namely "Indirect" ELISA , Sandwich ELISA , Competitive ELISA and ELISA Reverse method & device (ELISA-R m&d).
The indirect ELISA is used primarily to determine the strength and/or amount of antibody response in a sample, whether it is from the serum of an immunized animal or the cell supernatant from growing hybridoma clones.
The Sandwich ELISA measures the amount of antigen between two layers of antibodies (i.e. capture and detection antibody). The antigen to be measured must contain at least two antigenic sites capable of binding to antibody, since at least two antibodies act in the sandwich.
A competitive ELISA provides an alternative to the direct ELISA and can measure the amount of antigen in a sample, however it is much less sensitive.
Competitive ELISA:

Last but not least, Reverse method & device (ELISA-R m&d). it is a newer technique that uses an solid phase made up of an immunosorbent polystyrene rod with 4-12 protruding ogives. The entire device is immersed in a test tube containing the collected sample and the following steps (washing, incubation in conjugate and incubation in chromogenous) are carried out by dipping the ogives in microwells of standard microplates pre-filled with reagents.
In summary, ELISA is:
• Enzyme-linked immunosorbant assat
• An antibody to detect antigen
• An antibody labeled with indicator (e.g. horse radish peroxidase)
• A colour reaction
• A system that requires one molecule to be attached to a solid surface
An ELISA plate:
 Molecular Virology
Molecular Virology
Molecular Virology is the study of viruses at the molecular level. In particular, this includes the analysis of individual viral genes and gene products, and their interaction with host (human, plant or animal) cellular proteins. In short: molecular virology is:
• Study genome organization
Genome organization refers to the sequential, not the structural organization of the genome
• Expression of viral genome
Different viruses have different strains of genomes that have been subdivided into different serotypes. The viral genome structures of these three serotypes were studied.
• Replication of viral genome and Progeny virus
• Molecular basis of viral replication
- Attachment and entry
- Replication and translation
- Assembly or virions
- Virus-host interactions
Viral replication. During the process of viral replication, a virus induces a living host cell to synthesize the essential components for the synthesis of new viral particles. The particles are then assembled into the correct structure, and the newly formed virions escape from the cell to infect other cells.
The first step in the replication process is attachment. In this step, the virus adsorbs to a susceptible host cell. High specificity exists between virus and cell, and the envelope spikes may unite with cell surface receptors. Receptors may exist on bacterial pili or flagella or on the host cell membrane.
The next step is penetration of the virus or the viral genome into the cell. This step may occur by phagocytosis; or the envelope of the virus may blend with the cell membrane; or the virus may “inject” its genome into the host cell. The latter situation occurs with the bacteriophage when the tail of the phage unites with the bacterial cell wall and enzymes open a hole in the wall. The DNA of the phage penetrates through this hole.
The replication steps of the process occur next. The protein capsid is stripped away from the genome, and the genome is freed in the cell cytoplasm. If the genome consists of RNA, the genome acts as a messenger RNA molecule and provides the genetic codes for the synthesis of enzymes. The enzymes are used for the synthesis of viral genomes and capsomeres and the assembly of these components into new viruses. If the viral genome consists of DNA, it provides the genetic code for the synthesis of messenger RNA molecules, and the process proceeds.
Molecular Biology and Others
• Analysis of viral proteins
- PAGE/ SDS-PAGE
SDS-PAGE, sodium dodecyl sulfate polyacrylamide gel electrophoresis, is a technique widely used in biochemistry, forensics, genetics and molecular biology to separate proteins according to their electrophoretic mobility (a function of length of polypeptide chain or molecular weight as well as higher order protein folding, posttranslational modifications and other factors).
- Western bolt
The western blot (alternatively, immunoblot) is an analytical technique used to detect specific proteins in a given sample of tissue homogenate or extract. It uses gel electrophoresis to separate native or denatured proteins by the length of the polypeptide (denaturing conditions) or by the 3-D structure of the protein (native/ non-denaturing conditions). The proteins are then transferred to a membrane (typically nitrocellulose or PVDF), where they are probed (detected) using antibodies specific to the target protein.
- Protein sequencing
Proteins are found in every cell and are essential to every biological process, protein structure is very complex: determining a protein's structure involves first protein sequencing - determining the amino acid sequences of its constituent peptides; and also determining what conformation it adopts and whether it is complexed with any non-peptide molecules.
Protein Sequencing:

- X-ray crystallography
X-ray crystallography is a method of determining the arrangement of atoms within a crystal, in which a beam of X-rays strikes a crystal and scatters into many different directions. From the angles and intensities of these scattered beams, a crystallographer can produce a three-dimensional picture of the density of electrons within the crystal. From this electron density, the mean positions of the atoms in the crystal can be determined, as well as their chemical bonds, their disorder and sundry other information.
• Analysis of viral genome
- Agarose geis
- Restriction analysis
- Sequencing
- Southern blot
A Southern blot is a method routinely used in molecular biology to check for the presence of a DNA sequence in a DNA sample. Southern blotting combines agarose gel electrophoresis for size separation of DNA with methods to transfer the size-separated DNA to a filter membrane for probe hybridization.
Southern Blot:

- Northern blot
The northern blot is a technique used in molecular biology research to study gene expression. It takes its name from its similarity to the Southern blot technique, named for biologist Edwin Southern.
Northern Blot:

- PCR/RT-PCR
PCR is Polymerase Chain Reaction (PCR) and RT-PCR is reverse transcriptase-PCR.
Southern Blot Analysis
A technique for separating DNA fragments by electrophoresis and then identifying a target fragment with a DNA probe. This method is named after its inventor, the British biologist Edwin M Southern in 1975. Specifically, Southern blot is a technique for transferring DNA of fragments from gel electrophoresis to nitrocellulose filter, specific fragments can then be detected by DNA probes, which were labelled radioactively or non-radioactively methods.
Northern blot
Northern blot is very similar to a Southern blot except that it is RNA rather than DNA which is extracted, run on a gel and transferred to a filter membrane. Both Northern and Southern Blot use electrophoresis and detection with a hybridization probe. The northern blot technique was developed in 1977 by James Alwine, David Kemp, and George Stark at Stanford University. In summary, Northern Blot is:
- Based on Southern Blot
- Analysis of RNA instead of DNA
- Named to reflect different molecule studied
Polymerase Chain Reaction
Polymerase chain reaction (PCR) enables researchers to produce millions of copies of a specific DNA sequence in approximately two hours. This automated process bypasses the need to use bacteria for amplifying DNA. In point form, Polymerase chain reaction (PCR):
• Is invented by Kary Mullis, 1193
• Named a Photocopier- Xeroxing DNA
• Generates million of copies of specific DNA in a few hours
• Quick and simple way of cloning genes in test tube
Steps of how PCR work
1. A PCR reaction starts with a denaturing step. Samples are heated to 94-96°C for one to several minutes to denature the target DNA so that they separate into single strands.
2. Next, the temperature is lowered to 50-65°C for one to several minutes. This allows the left and right primers to anneal to their complementary sequences. The primers are designed to bracket the DNA region to be amplified.
3. The temperature is raised to 72°C for one to several minutes. This allows the Taq polymerase to attach at each priming site and synthesize a new DNA strand.
4. The cycle repeats for about 20 to 30 times.
RT-PCR
Reverse transcriptase-PCR is the analysis of mRNA. It is a laboratory technique for amplifying a defined piece of a ribonucleic acid (RNA) molecule. The RNA strand is first reverse transcribed into its DNA complement or complementary DNA, followed by amplification of the resulting DNA using polymerase chain reaction. This can either be a 1 or 2 step process. Reverse transcription PCR is not to be confused with real-time polymerase chain reaction (Q-PCR) which is also sometimes wrongly abbreviated as RT-PCR. In summary RT-PCR:
• RNA is a single stranded, fragile, not stable
• Convert back to DNA called cDNA (‘Complementaty DNA’)
- The first step is first strand reaction
Complementary DNA (cDNA) is made from an mRNA template using dNTPs & reverse transcriptase. The components are combined with a DNA primer in a reverse transcriptase buffer for an hour at 37°C.
- The second step is second strand reaction
After the reverse transcriptase reaction is complete, cDNA has been generated from the original ss mRNA, standard PCR (called the “second strand reaction”) is initiated.
In the two-step RT, a thermostable DNA polymerase & the upstream and downstream DNA primers are added. Heating the reaction to temperatures above 37°C facilitates binding of DNA primers to the cDNA, & subsequent higher temperatures allow the DNA polymerase to make double-stranded DNA from the cDNA. Heating the reaction to 95°C melts the two DNA strands apart, enabling the primers to bind again at lower temperatures and begin the chain reaction again. After 30 cycles, millions of copies of the sequence of interest are generated.
Thats all for the Lecture 7 on Methods of Study of Viruses. Now, let’s proceed to Lecture 8.
---------------------------------------------------------------
LECTURE 8
Virus Host Interactions
Introduction to Virus
Viruses are strange things that straddle the fence between living and non-living. On the one hand, if they're floating around in the air or sitting on a doorknob, they're inert. They're about as alive as a rock. But if they come into contact with a suitable plant, animal or bacterial cell, they spring into action. They infect and take over the cell like pirates hijacking a ship.
A bacteriopahge:
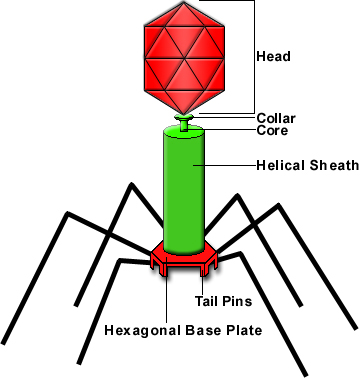 Morphology of Virus
Morphology of Virus
Viruses consist of two or three parts: all viruses have genes made from either DNA or RNA, long molecules that carry genetic information; all have a protein coat that protects these genes; and some have an envelope of fat that surrounds them when they are outside a cell. Viruses vary in shape from simple helical and icosahedral shapes, to more complex structures. They are about 100 times smaller than bacteria. The origins of viruses are unclear: some may have evolved from plasmids—pieces of DNA that can move between cells—others may have evolved from bacteria.
Genome of Virus
Viral genomes are circular, such as polyomaviruses, or linear, such as adenoviruses. The type of nucleic acid is irrelevant to the shape of the genome. Among RNA viruses, the genome is often divided up into separate parts within the virion and is called segmented. Each segment often codes for one protein and they are usually found together in one capsid. Every segment is not required to be in the same virion for the overall virus to be infectious, as demonstrated by the brome mosaic virus.
Viral Genes
• Immediate genes
- Transcription factors / enzymes
- Replication factors / enzymes
• Early genes
- Additional factors to regulate transcription, replication and translation
- Some structural proteins
• Late genes
- Mainly structural proteins
- Other proteins associated with virion / nucleocapsid
Viral Envelope
• Sources of envelope
- Nuclear membrane
- Endoplasmic reticulum
- Golgi body
- Plasma membrane
• Formation by budding off from membrane
Viral pathogenesis / infection
• Capacity of a virus to cause disease in a whole host
• Balance between viral replication, host defence and viral evasion of host defence
• Animal models only provide indication of viral pathogenesis in humans
• Complex
Sites of Viral Entry into Host
• Skin
- Cuts and abrasions.
• Conjunctiva (eyelids)
• Urogenital tract
• Respiratory tract
• Alimentary tract
Virus spread
• Systemic infection
- Many organs are infected
• Haematogenous spread
- Spread through the bloodstream
- Viremia
. Active / passive
. Primary / Secondary
• Neural spread
Virus shedding / Transmission
• Absolutely necessary for propagation / survival; exceptions
- Spread through germ cells
- Consumption of infected tissue (kuru, mad cow disease)
• Effective transmission depends on
- Virus concentration
- Route of transmission
Modes of Transmission
• Respiratory secretions
- Aerosols during speaking, sneezing, coughing, breathing, singing
- Virus easily inactivated by drying
- Further transmission via contaminated hands
- Rhinovirus, influenza, measles
• Saliva
- As for respiratory secretions
- Kissing
- Cytomegalovirus, herpesvirus, mumps, retroviruses
• Feaces
- Not relevant in developed countries
- Highly resistant to drying
- Enteric and hepatic viruses
• Blood
- Arthropod-borne viruses (Arboviruses)
. Flaviviruses, Togaviruses
- Direct blood / body fluids exposure
. Hepatitis HIV, haemorrhagic fevers (Bunyaviridae and Filoviridae)
. Healthcare workers at high risk
Virus-induced injury
• Cytopathic effect (CPE); cellular level
- Altered shape
- Detachment from substrate
- Lysis
- Membrane fusion; syncytium
- Membrane permeability
- Inclusion bodies
- Apoptosis
• Shutoff of cellular functions
- E.g. poliovirus shuts off cellular function in neurons resulting in cell death and hence paralysis
• Immunopathological lesions
- Impairment of immune response due to infection of immune cells (e.g. HIV on CD4+ and CD8+ T lymphocytes)
- Enhancement of immune response causing haemorrhagic fever (e.g. dengue, Hantaan, Ebola)
Stages in Virus Life Cycle
• Initiation of infection
- Attachment and entry
• Replication and expression
- Genome replication
- mRNA production, processing and translation
• Assembly and exit
• Viral pathogenesis
Virus Replication Cycle
1. Attachment
2. Penetration (entry)
3. Uncoating
4. Replication
5. Assembly
6. Release
Attachment
• Routes of host entry
- Endocytosis
; Non-specific endocytosis
; Phagocytosis
; Receptor-mediated endocytosis
- Fusion
- Direct penetration
• Specific attachment of virus attachment protein (VAP) to host cell receptor
• Receptor mediates viral entry into cell
• Coreceptors
Attachment is a specific binding between viral capsid proteins and specific receptors on the host cellular surface. This specificity determines the host range of a virus. For example, HIV infects only human T cells, because its surface protein, gp120, can interact with CD4 and receptors on the T cell's surface. This mechanism has evolved to favor those viruses that only infect cells in which they are capable of replication. Attachment to the receptor can induce the viral-envelope protein to undergo changes that results in the fusion of viral and cellular membranes.
Receptors
• Any surface molecule of the cell
- Glycoprotein
- Glycolipid
• Multiple receptors
• Host range and tissue tropism
Penetration (entry)
Penetration follows attachment; viruses enter the host cell through receptor mediated endocytosis or membrane fusion. This is often called viral entry. The infection of plant cells is different to that of animal cells. Plants have a rigid cell wall made of cellulose and viruses can only get inside the cells following trauma to the cell wall. Viruses such as tobacco mosaic virus can also move directly in plants, from cell-to-cell, through pores called plasmodesmata. Bacteria, like plants, have strong cell walls which a virus must breach to infect the cell. Some viruses have evolved mechanisms which inject their genome into the bacterial cell while the viral capsid remains outside.
Uncoating
• Virus genome is sensitive to external environment:
- Mechanical shearing
- UV radiation
- pH
- Dehydration
- Enzymes
• Capsid and envelope provide protection
• Successful / productive infection
- Introduction of viral genome into cell
- Susceptible
- Permissive
- Tissue tropism
Uncoating is a general term for the events which occur after penetration, in which the capsid is removed and the virus genome exposed, usually in the form of a nucleoprotein complex. Uncoating is a process in which the viral capsid is degraded by viral enzymes or host enzymes thus releasing the viral genomic nucleic acid.
Fusion
• From endosome
- Mediated by fall in pH
- Fusion from within
• Fusion from without (with plasma membrane)
- Attachment followed by fusion mediated by fusion protein
Direct penetration
• Translocation of entire virus particle
• Mediated by receptor
• Least understood mechanism of entry
Replication
• Genome replication and gene expression very closely linked
• Characteristics depends on nature of genome
Replication involves synthesis of viral messenger RNA (mRNA) for viruses except positive sense RNA viruses (see above), viral protein synthesis and assembly of viral proteins and viral genome replication. The replication strategy of the virus depends on the nature of its genome. Viruses can be classified into seven (arbitrary) groups:
I: Double-stranded DNA (Adenoviruses; Herpesviruses; Poxviruses, etc)
II: Single-stranded (+)sense DNA (Parvoviruses)
III: Double-stranded RNA (Reoviruses; Birnaviruses)
IV: Single-stranded (+)sense RNA (Picornaviruses; Togaviruses, etc)
V: Single-stranded (-)sense RNA (Orthomyxoviruses, Rhabdoviruses, etc)
VI: Single-stranded (+)sense RNA with DNA intermediate in life-cycle (Retroviruses)
VII: Double-stranded DNA with RNA intermediate (Hepadnaviruses)
How a virus cell replicates:
 Assembly
Assembly
• Assembly depends on site of synthesis
• Sites of protein synthesis and processing
- Endoplasmic reticulum
- Golgi body
• Sites for assembly
- Nucleus
- Endoplasmic reticulum
- Golgi body
Following the assembly of the virus particles, post-translational modification of the viral proteins often occurs. In viruses such as HIV, this modification, (sometimes called maturation), occurs after the virus has been released from the host cell.
Release
• Cell lysis
• Budding at plasma membrane
• Accumulation of particles in vesicles and release via exocytosis
Viruses are released from the host cell by lysis—a process that kills the cell by bursting its membrane. Enveloped viruses (e.g., HIV) typically are released from the host cell by budding. During this process, the virus acquires its envelope which is derived from the host's cell membrane.
Virus growth curve
• Eclipse / Latent stage
- Fall in virus titre when no infectious particles present during the replication process
- Virus not detected in external medium until released
• Virus burst
- New progeny virus assembled and released
Baltimore Classification
Baltimore is the classification base on genome, how they replicate and the relationship of virus with the mRNA. As mentioned in the replication section above, Baltimore classification has 7 classes.
Class I: Double-stranded DNA
• Two groups:
- Replication exclusively nuclear; very dependent on host cell factors
- Replication in cytoplasm; viral genome contains all factors for genome replication and transcription (Poxviridae)
Class II: Single-stranded DNA
• Replication of genome in nucleus
• Double-stranded DNA formed to make new single-stranded daughters
• Extreme parasitism
- Dependent on super-infecting viruses like adenovirus or herpesvirus for early regulatory genes
-
Chemical or UV light that induce similar virus-induced changes to host cell
• Parvoviridae
Class III: Double-stranded RNA
• Genome in several fragments
• Replication, transcription, translation regulated separately
• Monocistronic mRNA (code for 1 protein)
• All activity in cytoplasm; no nucleus
Class IV: Single-stranded Positive RNA
• Majority of animal and plant viruses
• Group 1
- Polycistronic mRNA
- Polyprotein formed and cleaved
• Group 2
- Complex transcription process
- 2 rounds of translation before formation of genomic RNA
Class V: Single-stranded Negative RNA
• Group 1:
- Non-segmented genome
- Transcription of negative RNA by RNA-dependent RNA polymerase to give monocistronic mRNA.
- mRNA also template for genome replication
- Ambisense organization
• Group 2:
- Orthomyxoviruses (segmented genome)
- Monocistronic mRNA in nucleus
- Virus transcriptase (in nucleocapsid)
Class VI: Single-stranded (+) sense RNA with DNA
• Retroviruses
• Single-stranded positive RNA, but with DNA intermediate
• Diploid
• Reverse transcription of viral RNA to dsDNA by viral RT
• Integration of dsDNA into host genome
• Viral RNA not used as mRNA
Group VII: Double-stranded DNA with RNA
• Double stranded DNA with RNA intermediate
• Not well understood
• Overlapping reading frames
• Hepadnaviridae
Regulation of Expression
• Transcriptional control
- Promoters on viral genome
- Early and late activators / enhancers
- Late repressors
- Viral transcriptases for RNA viruses not well understood
• Post-transcriptional control (RNA made)
- Splicing of polycistronic mRNA in nucleus
- Differential rate of splicing
- Control of mRNA from nucleus to cytoplasm
- Regulatory sequences found on introns
• Translational control
- Differential stability of mRNAs
- Secondary structures close to initiation sequence
- Problem due to overlapping reading frames
- IRES – internal ribosomal entry sites
- Frameshifting
- Pseudoknots
Thats all for Lecture 7 and 8. Lastly, lets go to Lecture 9 and 10 for our last virology reviews!